Uploading Files to the SPARC DAT-Core
Various ways to upload scientific files to the SPARC platform
Introduction
SPARC investigators are required to upload their scientific files to the DAT-Core. There are multiple ways for users to upload files to the platform. Here, we describe the various options and provide links to tutorials describing step-by-step instructions to upload files.
Step 1: Creating a dataset
All files belong to a dataset on the DAT-Core platform. A dataset contains both files and metadata and can be selectively shared with other users of the platform. You can create datasets through the web application or using one of the client tools.
Step 2: Uploading files
Once you have created a dataset, you can upload files to this dataset. You can create folders to separate files in a logical structure. After files are uploaded, you can associate files with metadata records. There are number of ways to upload files:
Upload files using the browser
Within a dataset, you can navigate to the Files tab. This is the main location where all files are organized.
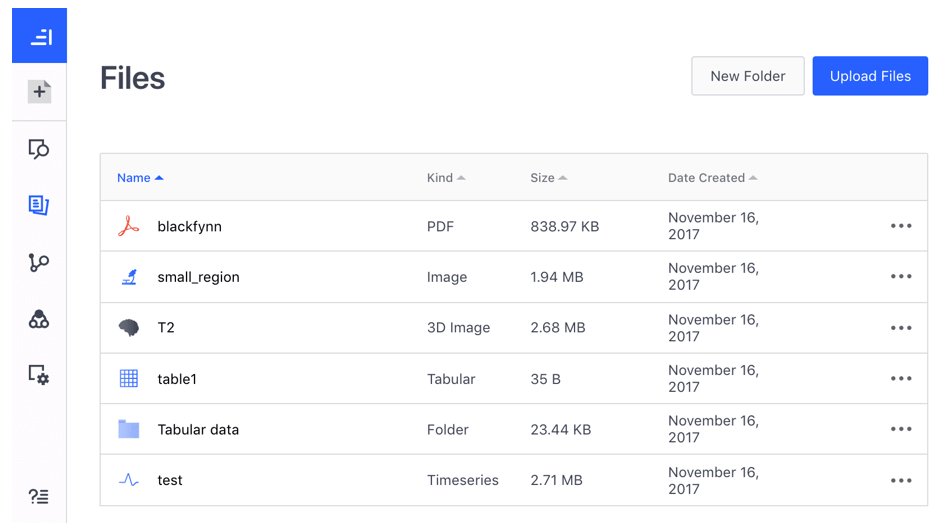
In the Files tab, you can create folders to organize your files on the platform. Click on the New Folder button to create a new folder. You can navigate into a folder by clicking on the folder in the file browser.
To upload files, you can drag files from your local machine directly into the file browser panel. A popup window will show the files that will be uploaded. Note that two of the files in the example below are combined as they belong to the same time series package.
Click Start Upload to begin uploading the files. During the upload process, you can safely hide the popup window as files will continue to be uploaded in the background. Once files are uploaded to the platform, they will show up in the file browser. If you drag a folder into the upload window, all nested folders will also be uploaded and the folder hierarchy will be maintained on the platform.
Depending on the file format, the platform will process the data to enable the data to be viewed in the browser-based data viewer. Processing of the data can take a while and you will be notified when this has completed.
For more information, see the file upload documentation for Pennsieve.
Upload folders/files using the Pennsieve CLI.
Pennsieve provides a Command Line Interface that can be used for uploading both small and very large datasets. Detailed information can be found here:
Upload files using the MBF Bioscience software
MBF Bioscience software can be used to upload files to the DAT-Core platform.
More details can be found here:
[Connecting MBF Bioscience software to the Pennsieve platform]
Upload files using Python
You can use the Open Source Pennsieve Python Library to upload files to the DAT-Core platform. In Python, you will first need to import the library and establish a connection to the platform. Then you can retrieve or create a dataset. The dataset object has a method to upload files to the dataset as the following example highlights. Note that uploading large files is currently not supported for Python; please use the Pennsieve Agent to programmatically upload large files. The Python client will rely on the Agent for uploading files in future releases.
# import Pennsieve
from pennsieve import Pennsieve
# create a client instance
ps = Pennsieve()
# create a dataset
ds = ps.create_dataset('Timeseries Dataset')
# upload data (3 files)
ds.upload('example_data/test.edf', 'example_data/testData.nev', 'example_data/testData.ns2')
More information can be found in the Pennsieve developers documentation for Python and the source code for the Python Library here.
Upload files using MATLAB
Finally, you can also upload files using the MATLAB client. The MATLAB client relies on the Pennsieve Agent to upload the files so you will have to both install the MATLAB client as well as the Pennsieve Agent. The upload command is a method of the PSDataset class and can be used as follows:
>> ps = Pennsieve();
>> ds = ps.datasets(1)
PSDataset with properties:
description: 'This is a dataset for the second tutorial.'
models: [1×3 PSModel]
items: [0×0 PSBaseDataNode]
name: 'tutorial2'
type: 'DataSet'
>> ds.upload('~/Downloads/example_data');A detailed tutorial on how to upload data using MATLAB can be found in the Pennsieve developers documentation for MATLAB and the MATLAB client source code.
Updated 10 months ago