Segmentation Viewer
The segmentation viewer provides a web preview of the full-resolution 3D morphological models generated from microscopy images.
To navigate through to the segmentation viewer:
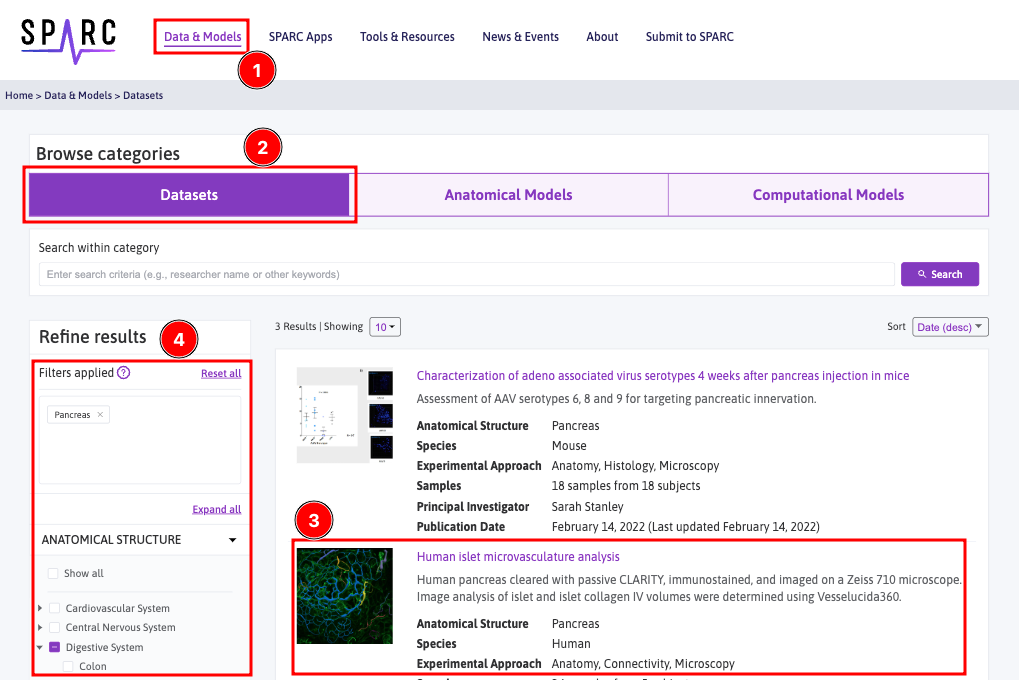
- Click Data & Models from the main menu.
- All the datasets are listed under the Datasets tab.
- Choose the dataset you want to explore by clicking on the dataset title. Example: Human Islet Microvasculature Analysis in Optically Cleared Pancreas using Vesselucida360 Analysis
- The data can be filtered if the expected data couldn't be found easily.
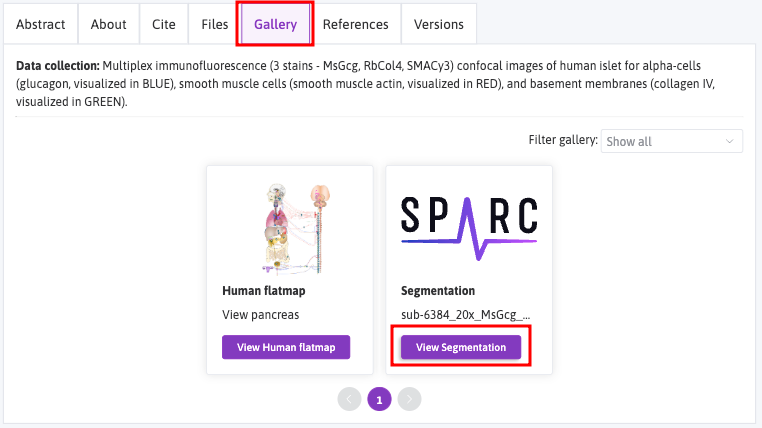
On the Dataset's page, select the Gallery tab to view thumbnails of the microscopy image data accompanied by segmentation, anatomical scaffolds, and flatmaps for particular datasets. Click on View Segmentation.
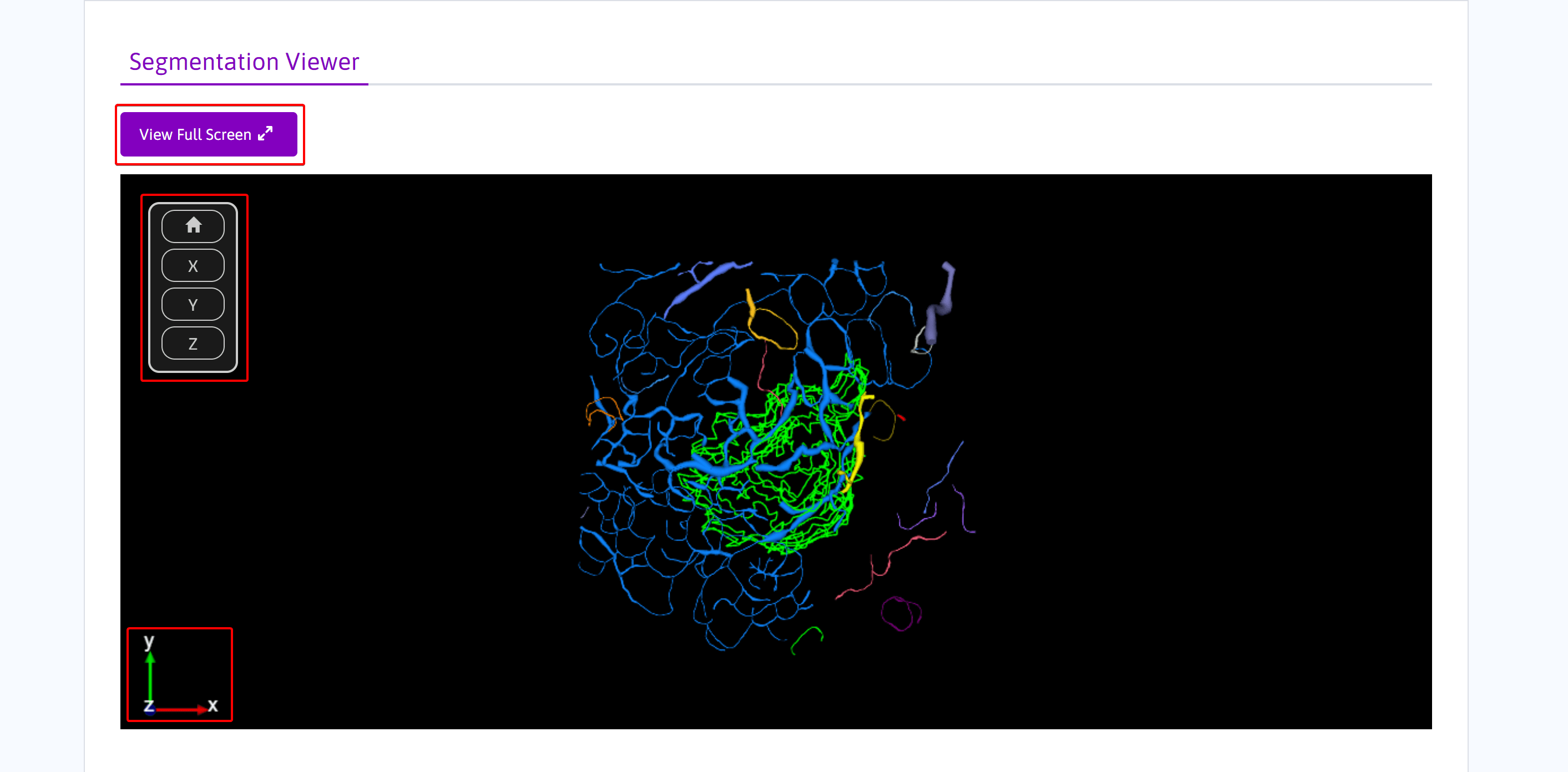
Use the following controls to interact with the segmentation viewer:
- Select View Full Screen to view the model in full screen in a new tab (optional).
- The Home icon resets the viewer to the default orientation and zoom level.
- A viewer aligned along the X points X-axis toward the viewer, the Y-axis points up, and the Z-axis points to the left.
- Similarly, the viewer alignment along the Y points Y-axis away from the viewer, the Z-axis points up, and the X-axis points right. The alignment of the viewer on the Z points Z-axis towards the viewer, the Y-axis points up, and the X-axis points right.
- Zoom in and zoom out of the model by scrolling with the mouse wheel.
- To rotate the model, click and drag with the mouse.
- To reposition the model, hold the shift key and drag with the mouse.
Below the segmentation viewer, file-level metadata is provided, including species, sex, age, and organ. The missing metadata fields may be present for datasets that have not been curated to the current SPARC Minimum Information Standard 3.0.
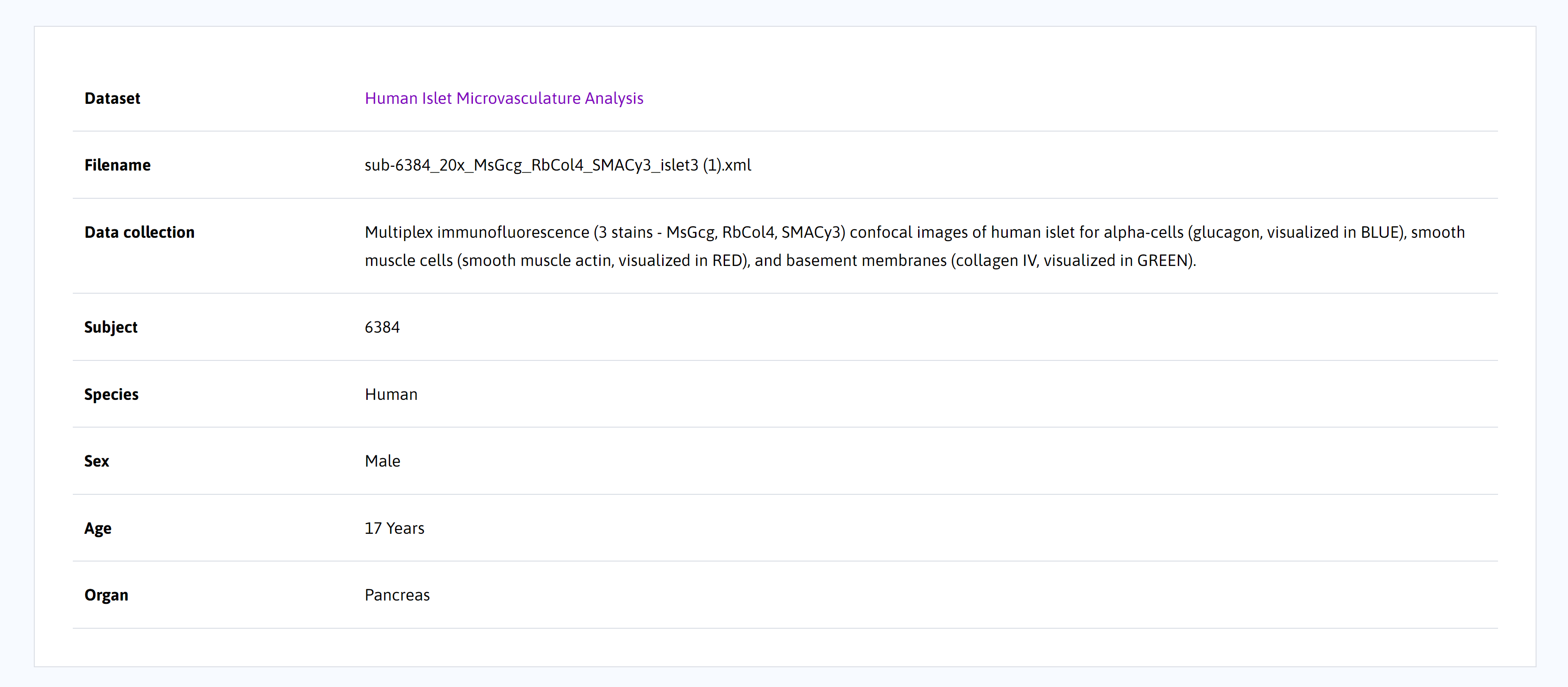
Note:The file-level metadata is pulled from the metadata within the segmentation file. This information is not curated and may be inconsistent with the curated data from the dataset.
Updated 11 months ago